Every data engineer must develop an understanding of the bias and variance tradeoff in machine learning (ML).
ML is used in more applications every day. Speech and image recognition, fraud detection, chatbots, and generative AI, are now commonplace uses of this technology. The growing use of ML is bringing the intricacies of how machine learning algorithms work out of specialized labs and into the mainstream of information technology.
Machine learning models cannot be a black box. Data engineers and users of large data sets must understand how to create and evaluate various algorithms and learning requirements when building and evaluating their ML models. Bias and variance in machine learning affect the overall accuracy of any model (which can be interpreted from a confusion matrix), trust in its outputs and outcomes, and its capacity to train machines to learn.
In this article, we will discuss what bias and variance in machine learning are. We will also touch on how to deal with the bias and variance tradeoff in developing useful algorithms for your ML applications.
(New to ML? Read our ML vs AI explainer.)
Bias vs. variance, and the tradeoff
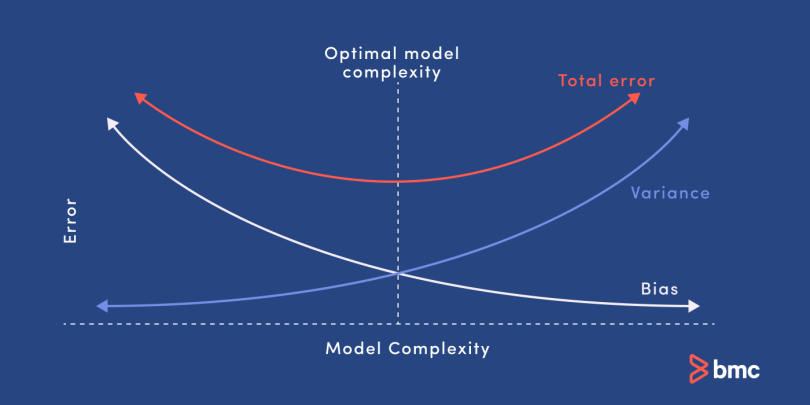
Bias and variance are two sources of error in predictive models. Getting the right balance between the bias and variance tradeoff is fundamental to effective machine learning algorithms. Here is a quick explanation of these concepts:
- Bias. Bias refers to error caused by a model for solving complex problems that is over simplified, makes significant assumptions, and misses important relationships in your data.
- Variance. Variance is an error caused by an algorithm that is too sensitive to fluctuations in data, creating an overly complex model that sees patterns in data that are actually just randomness.
- Bias–variance tradeoff. Minimizing errors caused by oversimplification and excessive complication requires finding the right balance or tradeoff between the two.
What is bias in machine learning?
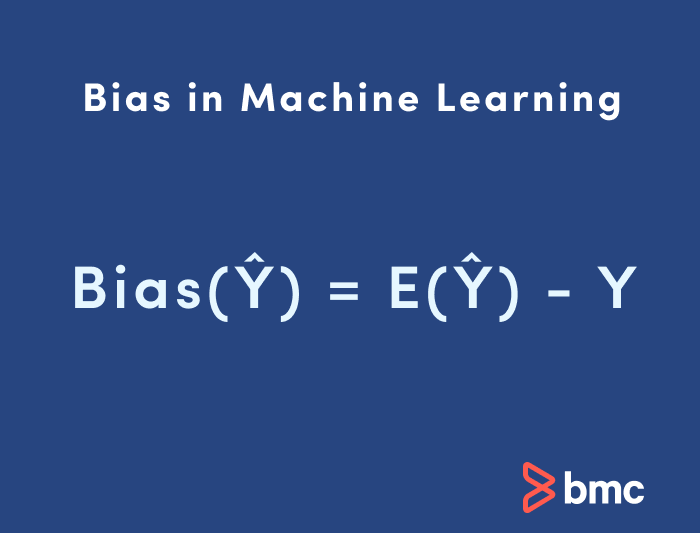

Bias in ML is sometimes called the “too simple” problem. Bias is considered a systematic error that occurs in the machine learning model itself due to incorrect assumptions in the ML process.
Technically, we can define bias as the error between average model prediction and the ground truth. Moreover, it describes how well the model matches the training data set:
- High bias. A model with a higher bias would not match the data set closely.
- Low bias. A low bias model will closely match the training data set.
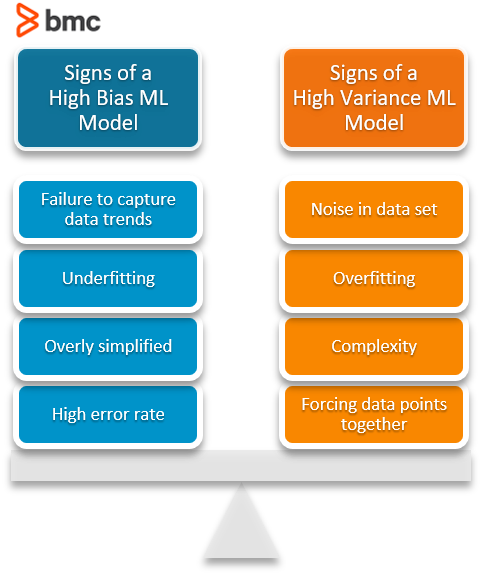
Characteristics of a high bias model include:
- Failure to capture proper data trends
- Potential towards underfitting
- More generalized/overly simplified
- High error rate
What is variance in machine learning?
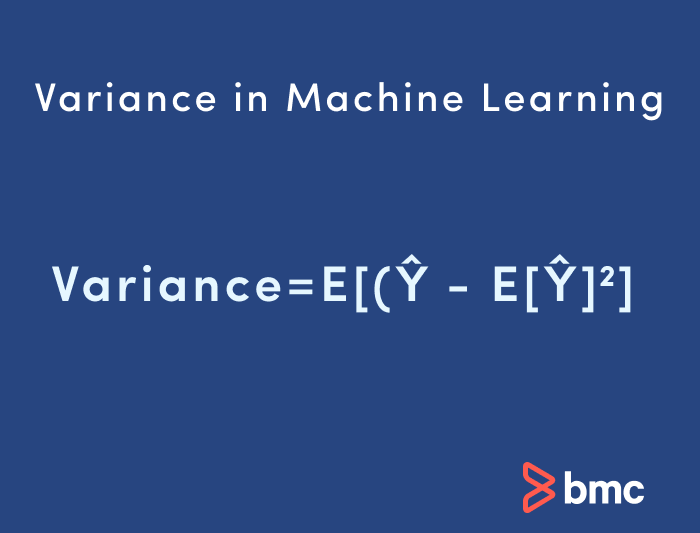

Variance in machine learning is sometimes called the “too sensitive” problem. Variance in ML refers to the changes in the model when using different portions of the training data set.
Simply stated, variance is the variability in the model prediction—how much the ML function can adjust depending on the given data set. Variance comes from highly complex models with a large number of features.
- Low variance. Models with high bias will have low variance.
- High variance. Models with high variance will have a low bias.
All these contribute to the flexibility of the model. For instance, a model that does not match a data set with a high bias will create an inflexible model with a low variance that results in a suboptimal machine learning model.
Characteristics of a high variance model include:
- Noise in the data set
- Potential towards overfitting
- Complex models
- Trying to put all data points as close as possible
Getting started with AIOps is easy. Learn how you can manage escalating IT complexity with ease! ›
Underfitting and overfitting
You can see from our fruit example how important it is that your model matches the data. How well your model “fits” the data directly correlates to how accurately it will perform in making identifications or predictions from a data set.
- Underfitting happens when your model is too simple to capture variations and patterns in your data. The machine doesn’t learn the right characteristics and relationships from the training data, and thus performs poorly with subsequent data sets. It might be trained on a red apple and mistake a red cherry for an apple.
- Overfitting happens when a model is too complex, with too much detail and random fluctuations or noise in the training data set. The machine erroneously sees this noise as true patterns, and thus is not able to generalize and see real patterns in subsequent data sets. It might be trained on many details of a specific type of apple and thus cannot find apples if they don’t have all these specific details.
Bias–variance trade-off
Bias and variance are inversely connected. It is impossible to have an ML model with a low bias and a low variance.
When a data engineer modifies the ML algorithm to better fit a given data set, it will lead to low bias—but it will increase variance. This way, the model will fit with the data set while increasing the chances of inaccurate predictions.
The same applies when creating a low variance model with a higher bias. While it will reduce the risk of inaccurate predictions, the model will not properly match the data set.
It’s a delicate balance between these bias and variance. Importantly, however, having a higher variance does not indicate a bad ML algorithm. Machine learning algorithms should be able to handle some variance.
We can tackle the trade-off in multiple ways…
- Increasing the complexity of the model to count for bias and variance, thus decreasing the overall bias while increasing the variance to an acceptable level. This aligns the model with the training dataset without incurring significant variance errors.
- Increasing the training data set can also help to balance this trade-off, to some extent. This is the preferred method when dealing with overfitting models. Furthermore, this allows users to increase the complexity without variance errors that pollute the model as with a large data set.
A large data set offers more data points for the algorithm to generalize data easily. However, the major issue with increasing the trading data set is that underfitting or low bias models are not that sensitive to the training data set. Therefore, increasing data is the preferred solution when it comes to dealing with high variance and high bias models.
This table lists common algorithms and their expected behavior regarding bias and variance:
| Algorithm | Bias | Variance |
| Linear Regression | High Bias | Less Variance |
| Decision Tree | Low Bias | High Variance |
| Bagging | Low Bias | High Variance (Less than Decision Tree) |
| Random Forest | Low Bias | High Variance (Less than Decision Tree and Bagging) |
Bias and variance calculation Python example
Let’s put these concepts into practice—we’ll calculate bias and variance using Python.
The simplest way to do this would be to use a library called mlxtend (machine learning extension), which is targeted for data science tasks. This library offers a function called bias_variance_decomp that we can use to calculate bias and variance.
We will be using the Iris data data set included in mlxtend as the base data set and carry out the bias_variance_decomp using two algorithms: Decision Tree and Bagging.
Decision tree example
from mlxtend.evaluate import bias_variance_decomp
from sklearn.tree import DecisionTreeClassifier
from mlxtend.data import iris_data
from sklearn.model_selection import train_test_split
# Get Data Set
X, y = iris_data()
X_train_ds, X_test_ds, y_train_ds, y_test_ds = train_test_split(X, y,
test_size=0.3,
random_state=123,
shuffle=True,
stratify=y)
# Define Algorithm
tree = DecisionTreeClassifier(random_state=123)
# Get Bias and Variance - bias_variance_decomp function
avg_expected_loss, avg_bias, avg_var = bias_variance_decomp(
tree, X_train_ds, y_train_ds, X_test_ds, y_test_ds,
loss='0-1_loss',
random_seed=123,
num_rounds=1000)
# Display Bias and Variance
print(f'Average Expected Loss: {round(avg_expected_loss, 4)}n')
print(f'Average Bias: {round(avg_bias, 4)}')
print(f'Average Variance: {round(avg_var, 4)}')
Result:

Bagging example
from mlxtend.evaluate import bias_variance_decomp
from sklearn.tree import DecisionTreeClassifier
from mlxtend.data import iris_data
from sklearn.model_selection import train_test_split
from sklearn.ensemble import BaggingClassifier
# Get Data Set
X, y = iris_data()
X_train_ds, X_test_ds, y_train_ds, y_test_ds = train_test_split(X, y,
test_size=0.3,
random_state=123,
shuffle=True,
stratify=y)
# Define Algorithm
tree = DecisionTreeClassifier(random_state=123)
bag = BaggingClassifier(base_estimator=tree,
n_estimators=100,
random_state=123)
# Get Bias and Variance - bias_variance_decomp function
avg_expected_loss, avg_bias, avg_var = bias_variance_decomp(
bag, X_train_ds, y_train_ds, X_test_ds, y_test_ds,
loss='0-1_loss',
random_seed=123,
num_rounds=1000)
# Display Bias and Variance
print(f'Average Expected Loss: {round(avg_expected_loss, 4)}n')
print(f'Average Bias: {round(avg_bias, 4)}')
print(f'Average Variance: {round(avg_var, 4)}')
Result:
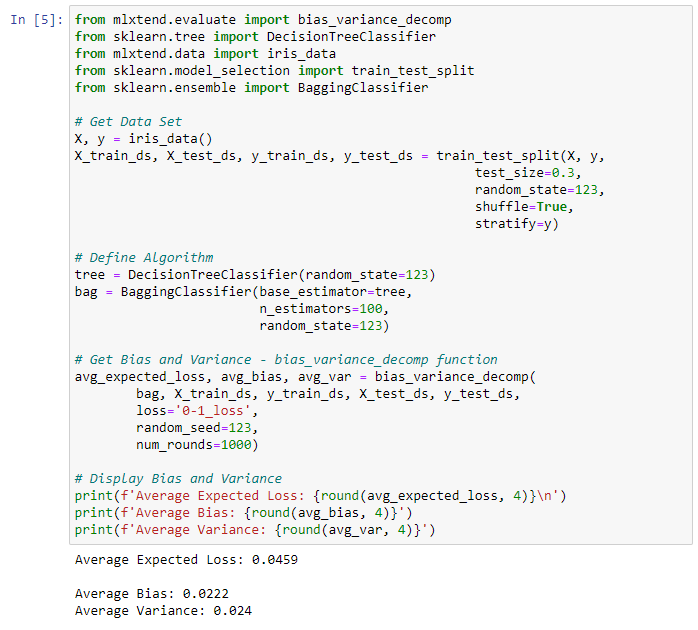
Each of the above functions will run 1,000 rounds (num_rounds=1000) before calculating the average bias and variance values. We can reduce the variance without affecting bias, using a bagging classifier. The higher the algorithm complexity, the lower the variance.
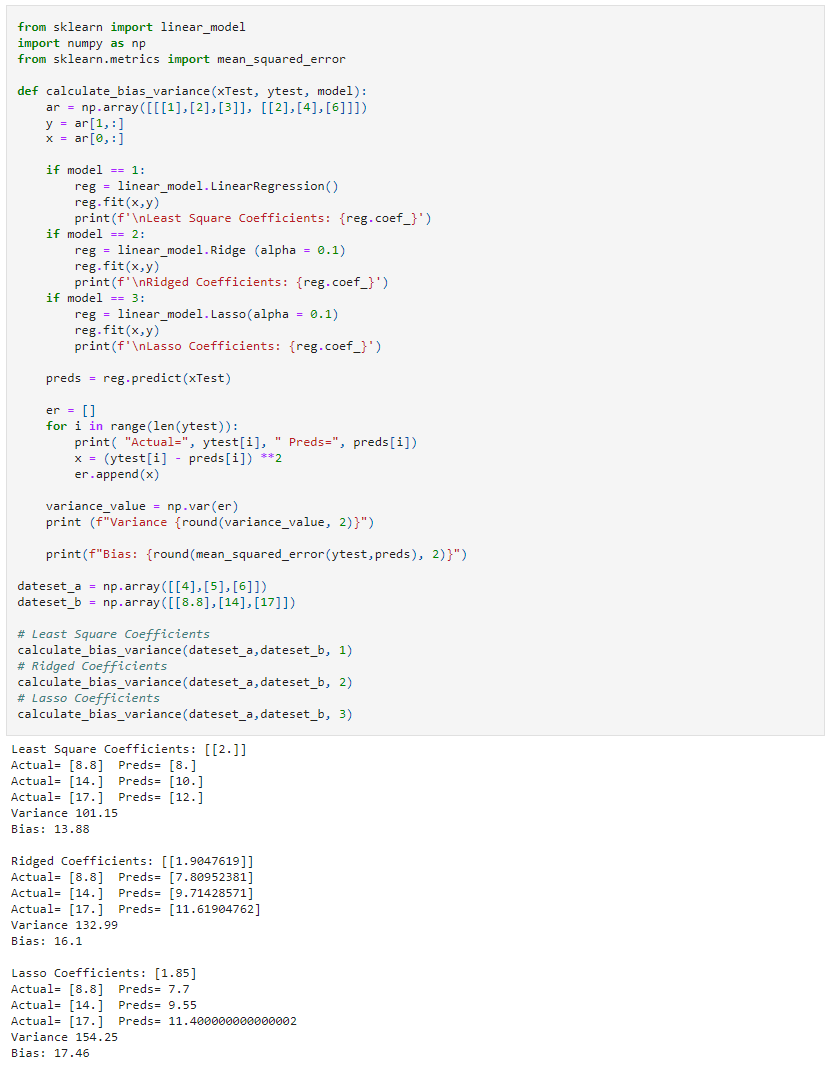
In the following example, we will look at three different linear regression models: least-squares, ridge, and lasso, using sklearn library. Since they are all linear regression algorithms, their main difference will be the coefficient value.
We can see those different algorithms lead to different outcomes in the ML process (bias and variance).
from sklearn import linear_model
import numpy as np
from sklearn.metrics import mean_squared_error
def calculate_bias_variance(xTest, ytest, model):
ar = np.array([[[1],[2],[3]], [[2],[4],[6]]])
y = ar[1,:]
x = ar[0,:]
if model == 1:
reg = linear_model.LinearRegression()
reg.fit(x,y)
print(f'nLeast Square Coefficients: {reg.coef_}')
if model == 2:
reg = linear_model.Ridge (alpha = 0.1)
reg.fit(x,y)
print(f'nRidged Coefficients: {reg.coef_}')
if model == 3:
reg = linear_model.Lasso(alpha = 0.1)
reg.fit(x,y)
print(f'nLasso Coefficients: {reg.coef_}')
preds = reg.predict(xTest)
er = []
for i in range(len(ytest)):
print( "Actual=", ytest[i], " Preds=", preds[i])
x = (ytest[i] - preds[i]) **2
er.append(x)
variance_value = np.var(er)
print (f"Variance {round(variance_value, 2)}")
print(f"Bias: {round(mean_squared_error(ytest,preds), 2)}")
dateset_a = np.array([[4],[5],[6]])
dateset_b = np.array([[8.8],[14],[17]])
# Least Square Coefficients
calculate_bias_variance(dateset_a,dateset_b, 1)
# Ridged Coefficients
calculate_bias_variance(dateset_a,dateset_b, 2)
# Lasso Coefficients
calculate_bias_variance(dateset_a,dateset_b, 3)
Result:
Considering bias and variance is crucial
Bias and variance are two key components that you must consider when developing any good, accurate machine learning model.
- Bias creates consistent errors in the ML model, which represents a simpler ML model that is not suitable for a specific requirement.
- On the other hand, variance creates variance errors that lead to incorrect predictions seeing trends or data points that do not exist.
Users need to consider both these factors when creating an ML model. Generally, your goal is to keep bias as low as possible while introducing acceptable levels of variances. This can be done either by increasing the complexity or increasing the training data set.
In this balanced way, you can create an acceptable machine learning model.
Related reading
- BMC Machine Learning & Big Data Blog
- Supervised, Unsupervised & Other Machine Learning Methods
- Anomaly Detection with Machine Learning: An Introduction
- Top Machine Learning Architectures Explained
- Machine Learning Frameworks To Use Today
- Data Ethics for Companies
These postings are my own and do not necessarily represent BMC's position, strategies, or opinion.
See an error or have a suggestion? Please let us know by emailing [email protected].